Engineered Single Amino Acid Substitutions Protect Hematopoietic Stem and Progenitor Cells From CD123 Targeted Immunotherapy
E. Landmann1,2*, A. Devaux1,2*, R. Lepore1,2,3,4*, R. Marone1,2*, C. Engdahl1,2, M. Hasiuk1,2, G. Capoferri1,2, A. Wiederkehr4, L. Wellinger4, A. Sinopoli4, V. Do Sacramento3, A. Haydn4, L. Garcia Prat3, C. Divsalar4, A. Camus3, J. Brault4, L. Xu5, T. Schwede6, M.H. Porteus7, S. Urlinger3,4, L.T. Jeker1,2,3
*These authors contributed equally
1 Transplantation Immunology & Nephrology, University Hospital Basel, Basel, Switzerland; 2 Department of Biomedicine, University of Basel, Basel, Switzerland; 3 CimeioTherapeutics AG, Basel, Switzerland; 4 Ridgeline Discovery GmbH, Basel, Switzerland; 5 Department of Pathology, Stanford University, Palo Alto, CA, USA; 6 Biozentrum, University of Basel, Basel, Switzerland; 7 Department of Pediatrics, Stanford University School of Medicine, Stanford, CA, USA
Introduction
- Targeted immunotherapy using monoclonal antibodies (mAb), T cell engager (TCE), or chimeric antigen receptor (CAR)) cells is standard of care in hematologic malignancies.
- CD123 (IL-3 receptor α-chain) is expressed in hematologic malignancies (e.g. acute myeloid leukaemia AML, blastic plasmacytoid dendritic cell neoplasm BPDCN), but also on healthy hematopoietic stem and progenitor cells (HSPCs)1.
- Shared antigen expression leads to on-target off-tumor effect with risk of extensive myelotoxicity following targeted cell depletion2.
- Proposed solution includes HSC transplantation with antigen-deleted HSPCs prior to immunotherapy3-5. This is, however, limited to dispensable targets and bears risk for antigen negative relapses.
Aim
Methods
- HEK cell lines stably expressing wildtype CD123 (HEK-CD123) and 28 CD123 variants with individual amino acid substitutions.
- Targeted immunotherapy:
- CSL362 biosimilar mAbMIRG123
- CSL362/OKT3 TCE
- Non-viral CRISPR/Cas9-based second generation CSL362-derived CAR T cells
- Non-viral engineering of HSPCs using CRISPR-Cas9 (High fidelity Cas9 RNPs and single strand oligodeoxynucleotides(ssODN) as homology directed repair template (HDRT).
- AAV6-mediated HDR editing of HSPCs (p-Stat5).
- In vivo: Injection of non-virally edited HSPCs into sublethally irradiated NSG mice.
Results
In silico design of human CD123 protein variants

a. Crystal structure of CD123 and the fragment antigen binding (Fab) of the CSL362 mAb(talacotuzumab). CSL362 epitope at the N-terminal domain (NTD) is indicated in light blue(red arrow). b. Per residue relative solvent accessibility computed on the CSL362-free (solid line) and -bound (dashed line) states based on the crystal structure. c.E51, S59 and R84 were identified as key amino acid (AA) residues for CSL362 binding and a total of 28 AA substitutions were chosen (not shown) that potentially disrupt binding but preserve function.
2. Abolished binding to theCSL362 biosimilar despite preserved expression of CD123
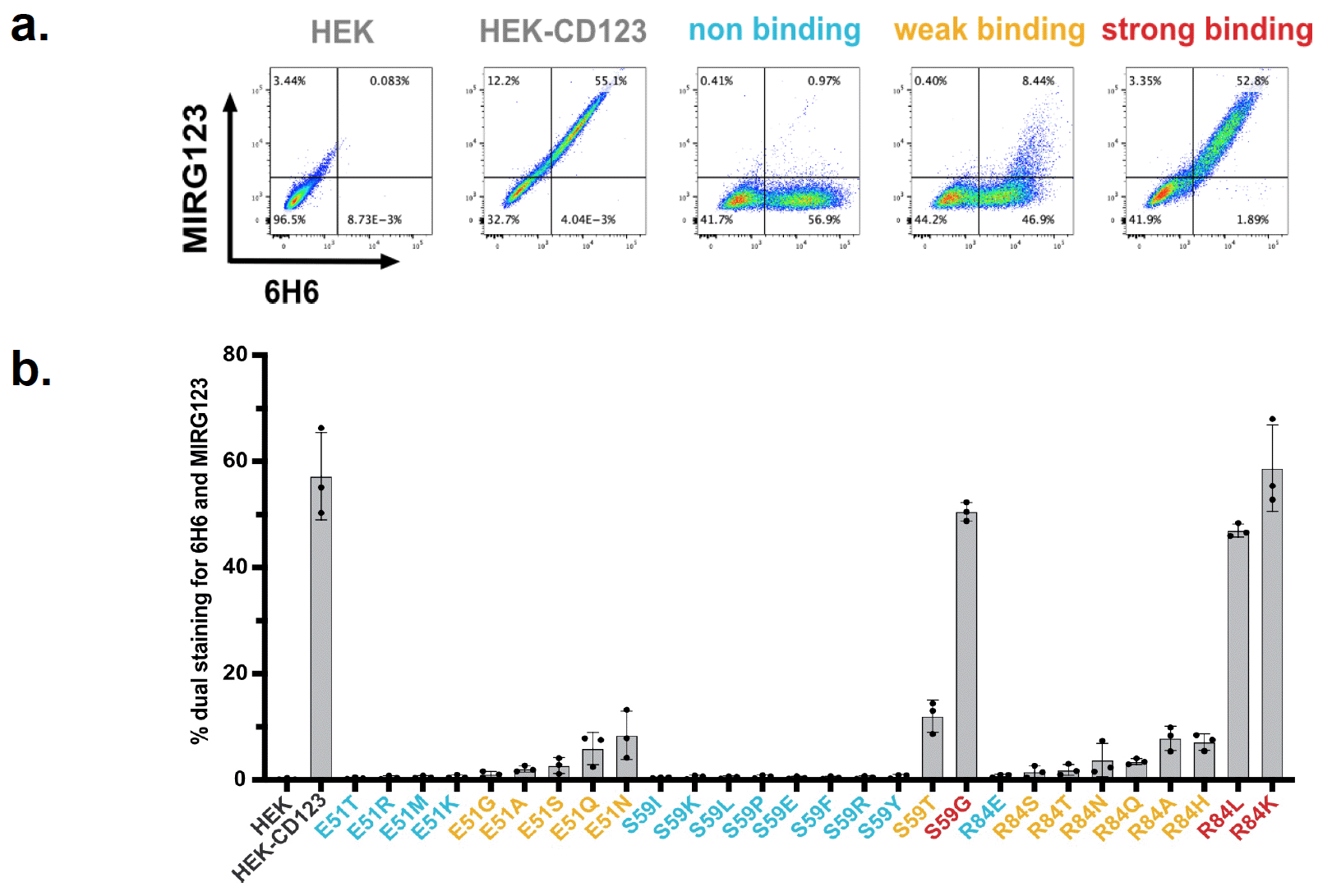
3. Biophysical characterization of selected CD123 variants

4. Engineered CD123 variants are shielded from TCE-and CAR-mediated cytotoxicity in vitro

5. Engineered human HSPCs expressing CD123 variants are functional, engraft and differentiate in vivo
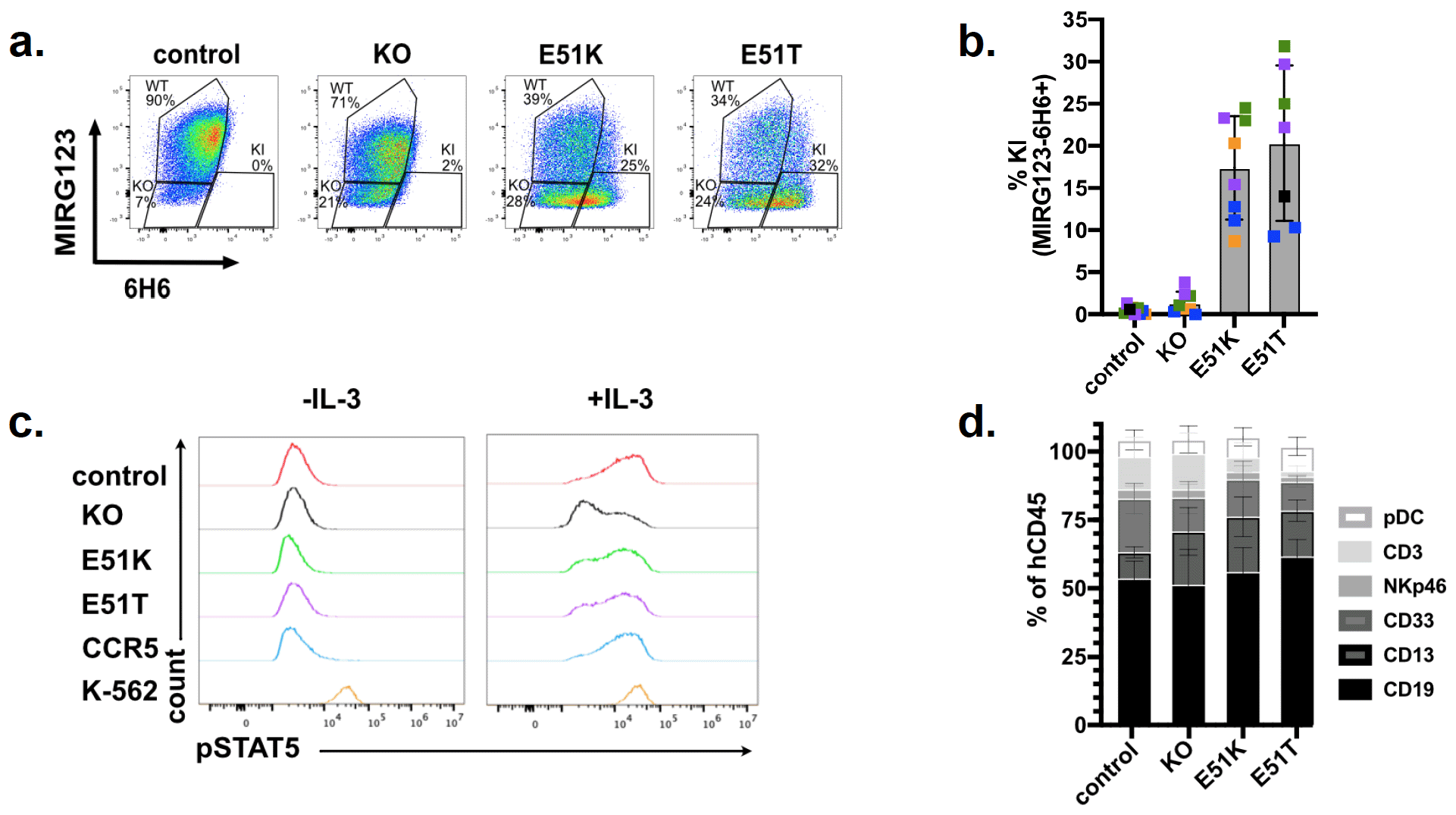
6. Engineered HSPCs enable tumor-selective CD123 immunotherapy
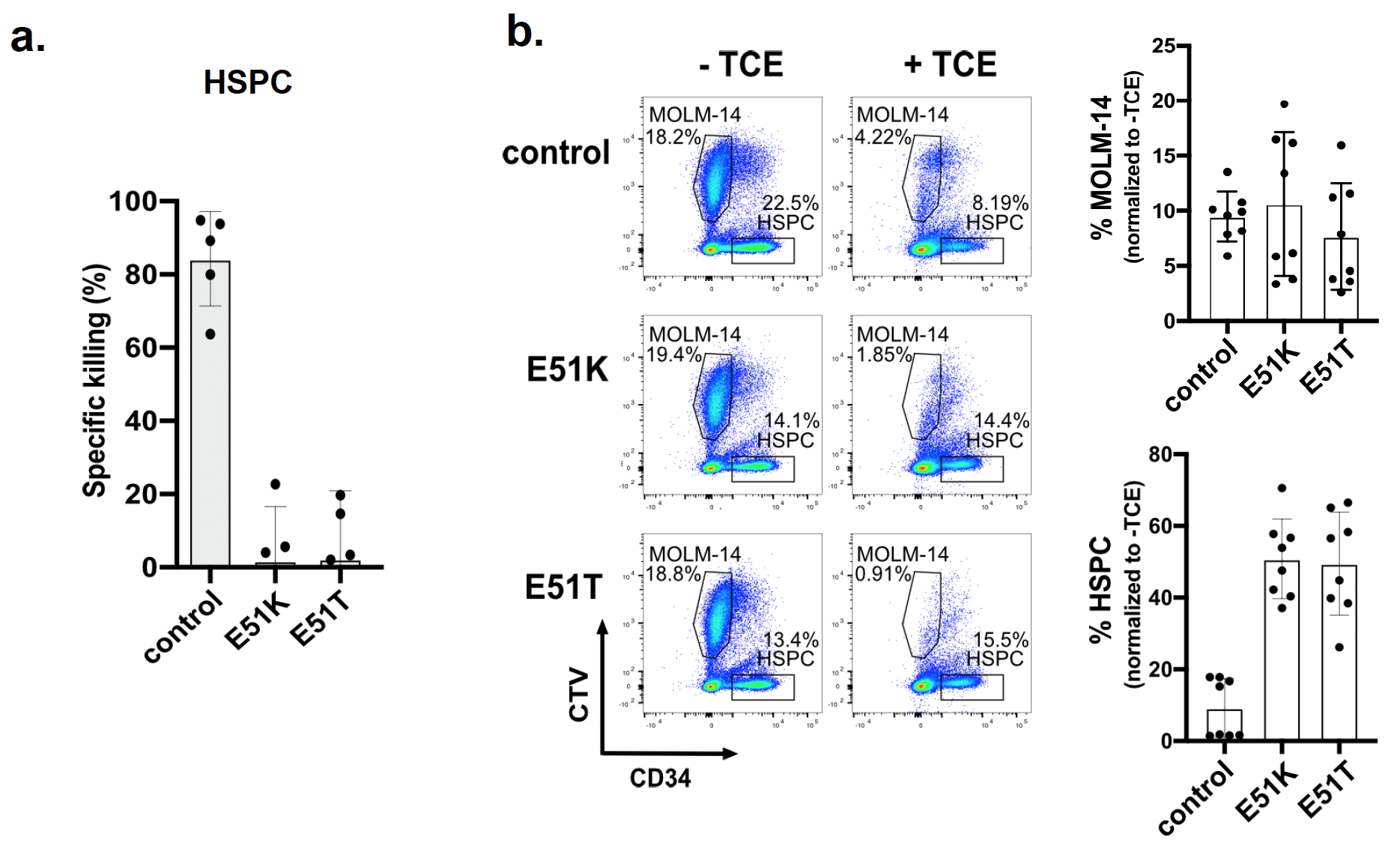
Conclusions
- Single amino acid substitutions abolish CSL362 binding but preserve function
- In vitro shielding of epitope engineered CD123 variants from TCE-and CAR-mediated killing
- Edited HSPCs are functional, engraft and differentiate regularly in vivo
- Tumor-selective killing of edited HSPCs in vitro
- Epitope engineering to shield cells may enable targeting of essential proteins
References
- Jordan, C. T. et al. The interleukin-3 receptor alpha chain is a unique marker for human acute myelogenous leukemia stem cells. Leukemia 14, 1777-1784, doi:10.1038/sj.leu.2401903 (2000)
- Gill, S. et al. Preclinical targeting of human acute myeloid leukemia and myeloablation using chimeric antigen receptor-modified T cells. Blood 123, 2343-2354, doi:10.1182/blood-2013-09-529537 (2014).
- Kim, M. Y. et al. Genetic Inactivation of CD33 in Hematopoietic Stem Cells to Enable CAR T Cell Immunotherapy for Acute Myeloid Leukemia. Cell 173, 1439-1453 e1419, doi:10.1016/j.cell.2018.05.013 (2018).
- Borot, F. et al. Gene-edited stem cells enable CD33-directed immune therapy for myeloid malignancies. Proc Natl AcadSciU S A 116, 11978-11987, doi:10.1073/pnas.1819992116 (2019)
- Humbert, O. et al. Engineering resistance to CD33-targeted immunotherapy in normal hematopoiesis by CRISPR/Cas9-deletion of CD33exon 2. Leukemia 33, 762-808, doi:10.1038/s41375-018-0277-8 (2019)
Acknowledgements
- Research Core Facilities (animal husbandry, flow cytometry) at the University of Basel and Department of Biomedicine.
- Dario Neriand MihaelaZavolanfor sharing reagents.
- Funding from European Research Council (ERC) European Union’s Horizon 2020 research and innovation programme (grant agreement No. 818806).
